9.3 Conjugate Priors and MAP Estimation#
Conjugate Priors#
Definition#
A prior distribution is conjugate to a likelihood if the posterior has the same distributional form as the prior.
Why Conjugacy Matters:
Closed-form posterior (no numerical integration needed)
Easy sequential updating
Interpretable hyperparameters
Computational efficiency
Common Conjugate Pairs#
Table of Conjugate Priors#
Likelihood |
Parameter |
Conjugate Prior |
Posterior |
|---|---|---|---|
Bernoulli(p) |
p |
Beta(α, β) |
Beta(α+k, β+n-k) |
Binomial(n, p) |
p |
Beta(α, β) |
Beta(α+k, β+n-k) |
Poisson(λ) |
λ |
Gamma(α, β) |
Gamma(α+Σx, β+n) |
Exponential(λ) |
λ |
Gamma(α, β) |
Gamma(α+n, β+Σx) |
Normal(μ, σ²) |
μ (σ² known) |
Normal(μ₀, σ₀²) |
Normal(…) |
Normal(μ, σ²) |
σ² (μ known) |
Inverse-Gamma |
Inverse-Gamma |
where:
k = number of successes
n = number of trials
Σx = sum of observations
Beta-Binomial (Detailed)#
The Setup#
Likelihood: Binomial
Prior: Beta
Posterior Derivation#
This is the kernel of:
Hyperparameter Interpretation#
Prior hyperparameters:
(\alpha) = “pseudo-counts” of successes
(\beta) = “pseudo-counts” of failures
(\alpha + \beta) = “effective sample size” of prior
Prior mean: (\frac{\alpha}{\alpha + \beta})
Posterior mean:
This is a weighted average of prior mean and MLE!
Python Implementation#
import numpy as np
import matplotlib.pyplot as plt
from scipy import stats
def beta_binomial_update(alpha_prior, beta_prior, k, n):
"""
Update Beta prior with Binomial data.
Parameters:
-----------
alpha_prior, beta_prior : Prior Beta parameters
k : Number of successes observed
n : Number of trials
Returns:
--------
alpha_post, beta_post : Posterior Beta parameters
"""
alpha_post = alpha_prior + k
beta_post = beta_prior + (n - k)
return alpha_post, beta_post
# Example
print("Beta-Binomial Conjugacy Example")
print("="*70)
# Prior
alpha_prior, beta_prior = 2, 2
print(f"\nPrior: Beta({alpha_prior}, {beta_prior})")
print(f"Prior mean: {alpha_prior/(alpha_prior + beta_prior):.3f}")
# Data
k, n = 15, 20 # 15 successes in 20 trials
print(f"\nData: {k} successes in {n} trials")
print(f"MLE: {k/n:.3f}")
# Posterior
alpha_post, beta_post = beta_binomial_update(alpha_prior, beta_prior, k, n)
print(f"\nPosterior: Beta({alpha_post}, {beta_post})")
print(f"Posterior mean: {alpha_post/(alpha_post + beta_post):.3f}")
# Visualize
p_range = np.linspace(0, 1, 1000)
prior = stats.beta(alpha_prior, beta_prior)
posterior = stats.beta(alpha_post, beta_post)
plt.figure(figsize=(10, 6))
plt.plot(p_range, prior.pdf(p_range), 'b--', linewidth=2, label='Prior', alpha=0.7)
plt.plot(p_range, posterior.pdf(p_range), 'r-', linewidth=2.5, label='Posterior')
plt.axvline(k/n, color='green', linestyle=':', linewidth=2, label=f'MLE = {k/n:.2f}')
plt.axvline(alpha_post/(alpha_post + beta_post), color='red', linestyle=':',
linewidth=1.5, label=f'Posterior mean = {alpha_post/(alpha_post + beta_post):.2f}')
plt.xlabel('p (probability)', fontsize=12)
plt.ylabel('Density', fontsize=12)
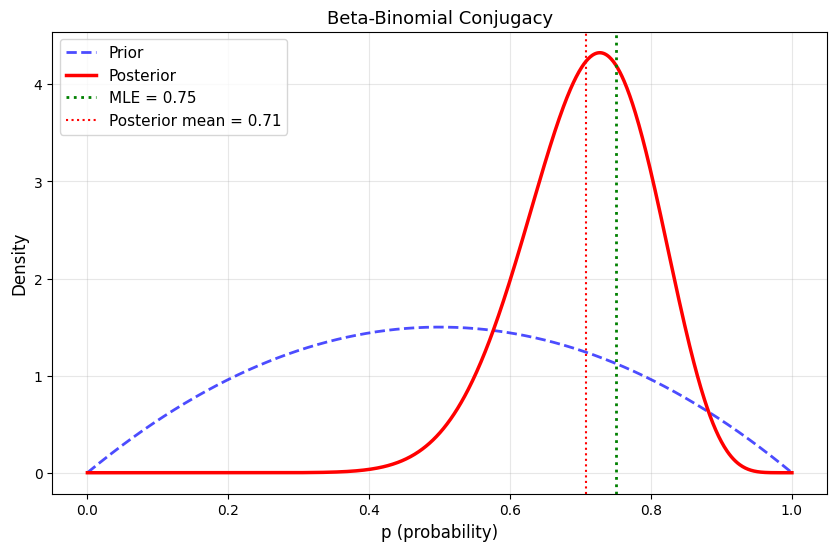
plt.title('Beta-Binomial Conjugacy', fontsize=13)
plt.legend(fontsize=11)
plt.grid(alpha=0.3)
plt.savefig('beta_binomial_conjugate.png', dpi=150, bbox_inches='tight')
plt.show()
Beta-Binomial Conjugacy Example
======================================================================
Prior: Beta(2, 2)
Prior mean: 0.500
Data: 15 successes in 20 trials
MLE: 0.750
Posterior: Beta(17, 7)
Posterior mean: 0.708
Gamma-Poisson (Detailed)#
The Setup#
Likelihood: Poisson observations (x_1, \ldots, x_n)
Prior: Gamma
Posterior Derivation#
This is the kernel of:
Hyperparameter Interpretation#
(\alpha) = prior “total count”
(\beta) = prior “observation periods”
Prior mean: (\frac{\alpha}{\beta})
Posterior mean: (\frac{\alpha + \sum x_i}{\beta + n})
Python Implementation#
import numpy as np
import matplotlib.pyplot as plt
from scipy import stats
np.random.seed(42)
# True parameter
true_lambda = 5.0
# Generate Poisson data
n = 30
data = np.random.poisson(true_lambda, n)
sum_x = np.sum(data)
print("Gamma-Poisson Conjugacy Example")
print("="*70)
print(f"\nTrue λ: {true_lambda}")
print(f"Data: {n} observations, sum = {sum_x}")
print(f"MLE: {sum_x/n:.3f}")
# Prior: Gamma(2, 1) - weakly informative
alpha_prior = 2
beta_prior = 1
print(f"\nPrior: Gamma({alpha_prior}, {beta_prior})")
print(f"Prior mean: {alpha_prior/beta_prior:.3f}")
# Posterior
alpha_post = alpha_prior + sum_x
beta_post = beta_prior + n
print(f"\nPosterior: Gamma({alpha_post}, {beta_post})")
print(f"Posterior mean: {alpha_post/beta_post:.3f}")
# Visualize
lambda_range = np.linspace(0, 10, 1000)
prior = stats.gamma(alpha_prior, scale=1/beta_prior)
posterior = stats.gamma(alpha_post, scale=1/beta_post)
plt.figure(figsize=(10, 6))
plt.plot(lambda_range, prior.pdf(lambda_range), 'b--', linewidth=2,
label='Prior', alpha=0.7)
plt.plot(lambda_range, posterior.pdf(lambda_range), 'r-', linewidth=2.5,
label='Posterior')
plt.axvline(true_lambda, color='black', linestyle='--', linewidth=1.5,
label=f'True λ = {true_lambda}', alpha=0.5)
plt.axvline(sum_x/n, color='green', linestyle=':', linewidth=2,
label=f'MLE = {sum_x/n:.2f}')
plt.axvline(alpha_post/beta_post, color='red', linestyle=':', linewidth=1.5,
label=f'Posterior mean = {alpha_post/beta_post:.2f}')
plt.xlabel('λ (rate parameter)', fontsize=12)
plt.ylabel('Density', fontsize=12)
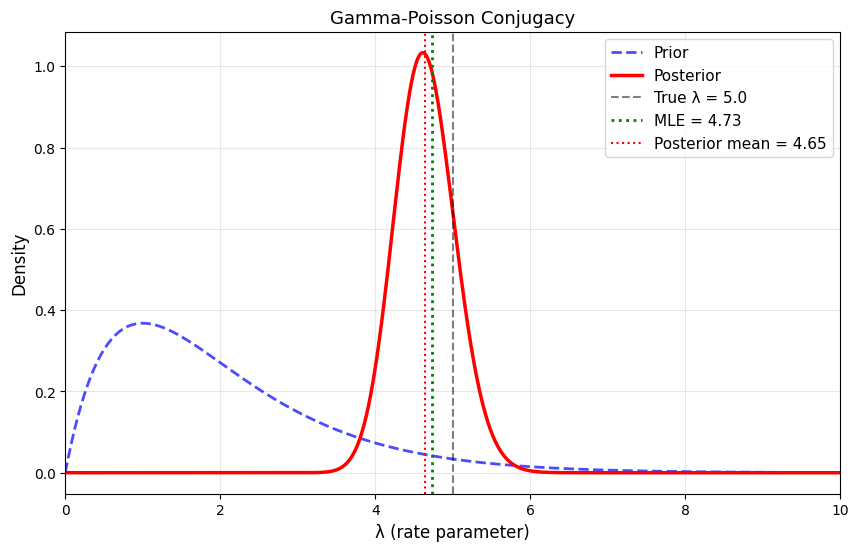
plt.title('Gamma-Poisson Conjugacy', fontsize=13)
plt.legend(fontsize=11)
plt.grid(alpha=0.3)
plt.xlim(0, 10)
plt.savefig('gamma_poisson_conjugate.png', dpi=150, bbox_inches='tight')
plt.show()
Gamma-Poisson Conjugacy Example
======================================================================
True λ: 5.0
Data: 30 observations, sum = 142
MLE: 4.733
Prior: Gamma(2, 1)
Prior mean: 2.000
Posterior: Gamma(144, 31)
Posterior mean: 4.645
Normal-Normal (Known Variance)#
The Setup#
Likelihood: (x_1, \ldots, x_n \sim N(\mu, \sigma^2)) where (\sigma^2) is known
Prior: (\mu \sim N(\mu_0, \sigma_0^2))
Posterior (Closed Form)#
The posterior is also Normal:
where:
Posterior mean:
Posterior variance:
(Precision of posterior = precision of prior + precision of data)
Interpretation#
Posterior mean is weighted average of prior mean and sample mean
Weights depend on relative precisions (inverse variances)
As (n \to \infty), posterior mean (\to \bar{x}) (data dominates)
Posterior variance always smaller than prior variance (we learned!)
Python Implementation#
import numpy as np
import matplotlib.pyplot as plt
from scipy import stats
np.random.seed(42)
# True parameter
true_mu = 5.0
sigma = 2.0 # Known variance
# Generate data
n = 10
data = np.random.normal(true_mu, sigma, n)
xbar = np.mean(data)
print("Normal-Normal Conjugacy Example")
print("="*70)
print(f"\nTrue μ: {true_mu}")
print(f"Known σ: {sigma}")
print(f"Data: n = {n}, x̄ = {xbar:.3f}")
print(f"MLE: {xbar:.3f}")
# Prior
mu_0 = 0.0
sigma_0 = 5.0
print(f"\nPrior: N({mu_0}, {sigma_0**2})")
# Posterior parameters
precision_prior = 1 / sigma_0**2
precision_data = n / sigma**2
precision_post = precision_prior + precision_data
sigma_n = np.sqrt(1 / precision_post)
mu_n = (precision_prior * mu_0 + precision_data * xbar) / precision_post
print(f"\nPosterior: N({mu_n:.3f}, {sigma_n**2:.3f})")
print(f"Posterior mean: {mu_n:.3f}")
print(f"Posterior std: {sigma_n:.3f}")
# Visualize
mu_range = np.linspace(-5, 10, 1000)
prior = stats.norm(mu_0, sigma_0)
posterior = stats.norm(mu_n, sigma_n)
plt.figure(figsize=(10, 6))
plt.plot(mu_range, prior.pdf(mu_range), 'b--', linewidth=2,
label=f'Prior: N({mu_0}, {sigma_0**2})', alpha=0.7)
plt.plot(mu_range, posterior.pdf(mu_range), 'r-', linewidth=2.5,
label=f'Posterior: N({mu_n:.1f}, {sigma_n**2:.1f})')
plt.axvline(true_mu, color='black', linestyle='--', linewidth=1.5,
label=f'True μ = {true_mu}', alpha=0.5)
plt.axvline(xbar, color='green', linestyle=':', linewidth=2,
label=f'MLE (x̄) = {xbar:.2f}')
plt.xlabel('μ', fontsize=12)
plt.ylabel('Density', fontsize=12)
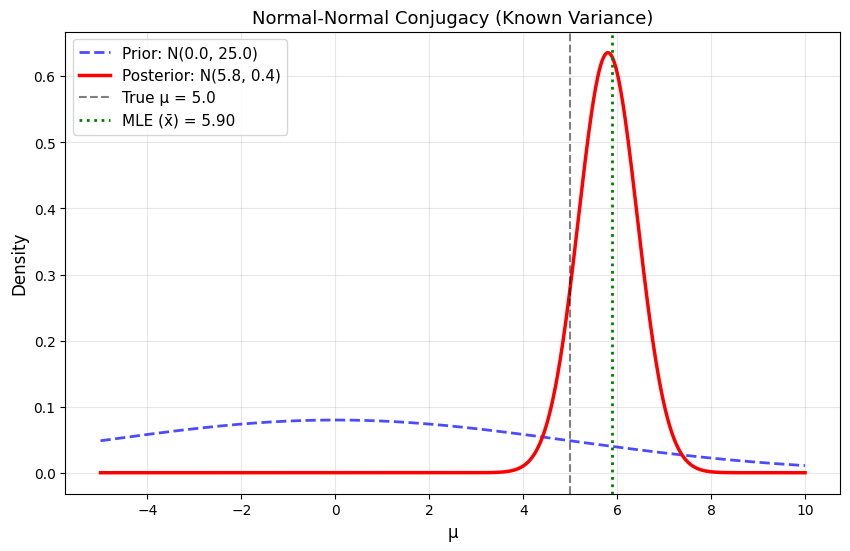
plt.title('Normal-Normal Conjugacy (Known Variance)', fontsize=13)
plt.legend(fontsize=11)
plt.grid(alpha=0.3)
plt.savefig('normal_normal_conjugate.png', dpi=150, bbox_inches='tight')
plt.show()
Normal-Normal Conjugacy Example
======================================================================
True μ: 5.0
Known σ: 2.0
Data: n = 10, x̄ = 5.896
MLE: 5.896
Prior: N(0.0, 25.0)
Posterior: N(5.803, 0.394)
Posterior mean: 5.803
Posterior std: 0.627
Maximum A Posteriori (MAP) Estimation#
Definition#
The MAP estimate is the mode of the posterior distribution:
Expanding:
Taking logs:
MAP vs. MLE#
MLE:
MAP:
Key insight: MAP = MLE + log prior penalty
MAP as Regularized MLE#
The log prior acts as a regularization term!
Example: Normal prior (\theta \sim N(0, \sigma_p^2))
So MAP becomes:
This is equivalent to L2 regularization (Ridge regression)!
When MAP = Posterior Mean#
For symmetric unimodal posteriors:
Mode = Mean = Median
MAP estimate = Posterior mean
Examples: Normal posterior, symmetric Beta
Python Example: Comparing Point Estimates#
import numpy as np
import matplotlib.pyplot as plt
from scipy import stats
from scipy.optimize import minimize_scalar
np.random.seed(42)
# Data
true_p = 0.7
n = 20
k = np.random.binomial(n, true_p)
print("Comparing Point Estimates: MLE, MAP, Posterior Mean")
print("="*70)
print(f"\nData: {k} heads in {n} flips")
print(f"True p: {true_p}")
# Prior: Beta(2, 2)
alpha_prior, beta_prior = 2, 2
# Posterior: Beta(alpha + k, beta + n - k)
alpha_post = alpha_prior + k
beta_post = beta_prior + (n - k)
print(f"\nPrior: Beta({alpha_prior}, {beta_prior})")
print(f"Posterior: Beta({alpha_post}, {beta_post})")
# Point estimates
p_mle = k / n
p_map = (alpha_post - 1) / (alpha_post + beta_post - 2) # Mode of Beta
p_mean = alpha_post / (alpha_post + beta_post) # Mean of Beta
p_median = stats.beta(alpha_post, beta_post).median()
print(f"\nPoint Estimates:")
print(f" MLE (no prior): {p_mle:.4f}")
print(f" MAP (mode): {p_map:.4f}")
print(f" Posterior mean: {p_mean:.4f}")
print(f" Posterior median: {p_median:.4f}")
print(f" True p: {true_p:.4f}")
# Visualize
p_range = np.linspace(0, 1, 1000)
posterior = stats.beta(alpha_post, beta_post)
plt.figure(figsize=(12, 6))
plt.plot(p_range, posterior.pdf(p_range), 'b-', linewidth=2.5,
label='Posterior')
plt.axvline(true_p, color='black', linestyle='--', linewidth=2,
label=f'True p = {true_p:.2f}', alpha=0.7)
plt.axvline(p_mle, color='green', linestyle=':', linewidth=2,
label=f'MLE = {p_mle:.3f}')
plt.axvline(p_map, color='red', linestyle='-.', linewidth=2,
label=f'MAP = {p_map:.3f}')
plt.axvline(p_mean, color='purple', linestyle=':', linewidth=2,
label=f'Post. mean = {p_mean:.3f}')
plt.xlabel('p', fontsize=12)
plt.ylabel('Posterior Density', fontsize=12)
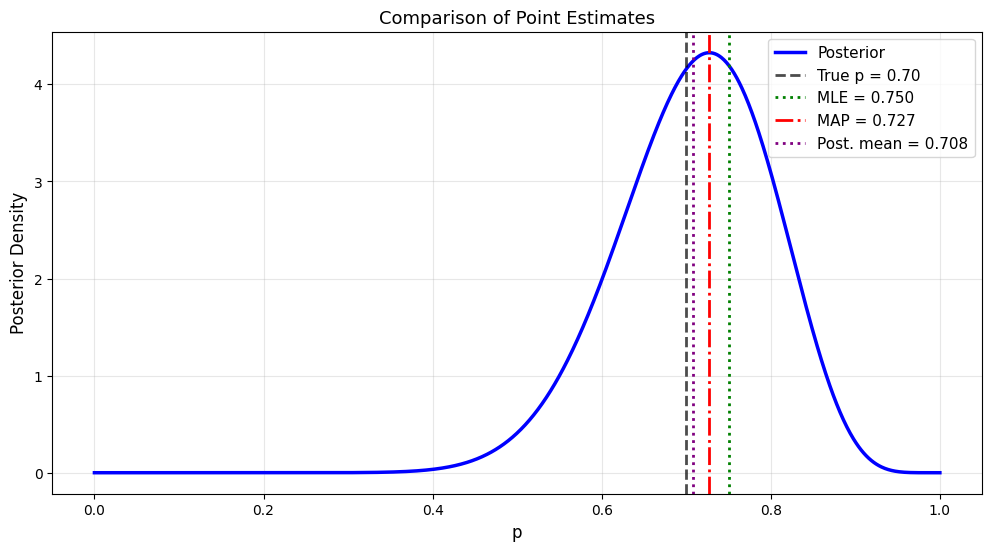
plt.title('Comparison of Point Estimates', fontsize=13)
plt.legend(fontsize=11)
plt.grid(alpha=0.3)
plt.savefig('point_estimates_comparison.png', dpi=150, bbox_inches='tight')
plt.show()
Comparing Point Estimates: MLE, MAP, Posterior Mean
======================================================================
Data: 15 heads in 20 flips
True p: 0.7
Prior: Beta(2, 2)
Posterior: Beta(17, 7)
Point Estimates:
MLE (no prior): 0.7500
MAP (mode): 0.7273
Posterior mean: 0.7083
Posterior median: 0.7142
True p: 0.7000
Summary#
Conjugate Priors#
Distribution |
Conjugate Prior |
Update Rule |
|---|---|---|
Bernoulli/Binomial |
Beta(α, β) |
α → α+k, β → β+n-k |
Poisson |
Gamma(α, β) |
α → α+Σx, β → β+n |
Normal (μ unknown) |
Normal(μ₀, σ₀²) |
Precision adds |
Point Estimates#
MLE: (\arg\max p(\text{data} \mid \theta))
MAP: (\arg\max p(\theta \mid \text{data})) = MLE + prior penalty
Posterior Mean: (\mathbb{E}[\theta \mid \text{data}]) = Optimal under squared loss
Key Insights#
✅ Conjugacy → closed-form posteriors ✅ MAP = regularized MLE ✅ Hyperparameters = “pseudo-data” ✅ As n → ∞, all estimates converge ✅ Prior matters most with small data