6.1 The Sample Mean#
The fundamental problem of statistical inference is: How can we learn about a population by examining a sample?
Populations and Samples#
Definitions#
Population: The entire collection of items we want to study.
Examples: All voters in a country, all manufactured chips, all possible coin flips
Sample: A subset of the population that we actually observe.
Examples: 1000 surveyed voters, 100 tested chips, 50 coin flips
Parameter: A numerical characteristic of the population (usually unknown).
Examples: Population mean μ, population variance σ²
Statistic: A numerical characteristic computed from a sample.
Examples: Sample mean x̄, sample variance s²
The Urn Model#
A useful mental model for sampling:
┌─────────────────────────────┐
│ Population (Urn) │
│ N items with values │
│ μ = population mean │
│ σ² = population variance │
└─────────────────────────────┘
↓ (draw n items)
┌─────────────────────────────┐
│ Sample │
│ n items: x₁, x₂, ..., xₙ │
│ x̄ = sample mean │
│ s² = sample variance │
└─────────────────────────────┘
The Sample Mean as an Estimator#
Definition#
Given a sample \((x_1, x_2, \ldots, x_n)\), the sample mean is:
Key Property: Unbiasedness#
Theorem: If \((X_1, X_2, \ldots, X_n)\) are independent random variables drawn from a distribution with mean μ, then:
Proof: $\( \begin{align} E[\bar{X}] &= E\left[\frac{1}{n}\sum_{i=1}^{n} X_i\right] \\ &= \frac{1}{n}\sum_{i=1}^{n} E[X_i] \\ &= \frac{1}{n}\sum_{i=1}^{n} \mu \\ &= \frac{1}{n} \cdot n\mu \\ &= \mu \end{align} \)$
This means the sample mean is an unbiased estimator of the population mean.
Variance of the Sample Mean#
The Key Result#
Theorem: If \((X_1, X_2, \ldots, X_n)\) are independent random variables from a distribution with variance σ², then:
Proof: $\( \begin{align} \text{Var}(\bar{X}) &= \text{Var}\left(\frac{1}{n}\sum_{i=1}^{n} X_i\right) \\ &= \frac{1}{n^2}\text{Var}\left(\sum_{i=1}^{n} X_i\right) \\ &= \frac{1}{n^2}\sum_{i=1}^{n} \text{Var}(X_i) \quad \text{(by independence)} \\ &= \frac{1}{n^2}\sum_{i=1}^{n} \sigma^2 \\ &= \frac{1}{n^2} \cdot n\sigma^2 \\ &= \frac{\sigma^2}{n} \end{align} \)$
Standard Error#
The standard error of the sample mean is:
Key Insight: The standard error decreases as (\sqrt{n}). To halve the standard error, you need 4 times as many samples!
Python Example: Sampling Distribution#
import numpy as np
import matplotlib.pyplot as plt
from scipy import stats
# Population: uniform distribution on [0, 10]
population_mean = 5.0
population_std = np.sqrt(100/12) # Var = (10-0)²/12
# Function to draw samples and compute sample mean
def sample_mean(n_samples, sample_size):
"""Draw n_samples, each of size sample_size, compute their means"""
means = []
for _ in range(n_samples):
sample = np.random.uniform(0, 10, sample_size)
means.append(np.mean(sample))
return np.array(means)
# Different sample sizes
sample_sizes = [5, 10, 30, 100]
n_samples = 10000
fig, axes = plt.subplots(2, 2, figsize=(12, 10))
axes = axes.ravel()
for idx, n in enumerate(sample_sizes):
# Generate sample means
sample_means = sample_mean(n_samples, n)
# Theoretical standard error
theoretical_se = population_std / np.sqrt(n)
# Empirical standard error
empirical_se = np.std(sample_means, ddof=1)
# Plot histogram
axes[idx].hist(sample_means, bins=50, density=True,
alpha=0.7, edgecolor='black', label='Empirical')
# Overlay theoretical normal distribution
x = np.linspace(sample_means.min(), sample_means.max(), 100)
axes[idx].plot(x, stats.norm.pdf(x, population_mean, theoretical_se),
'r-', linewidth=2, label='Theoretical N(5, σ/√n)')
axes[idx].axvline(population_mean, color='green', linestyle='--',
linewidth=2, label=f'True mean = {population_mean}')
axes[idx].set_xlabel('Sample Mean')
axes[idx].set_ylabel('Density')
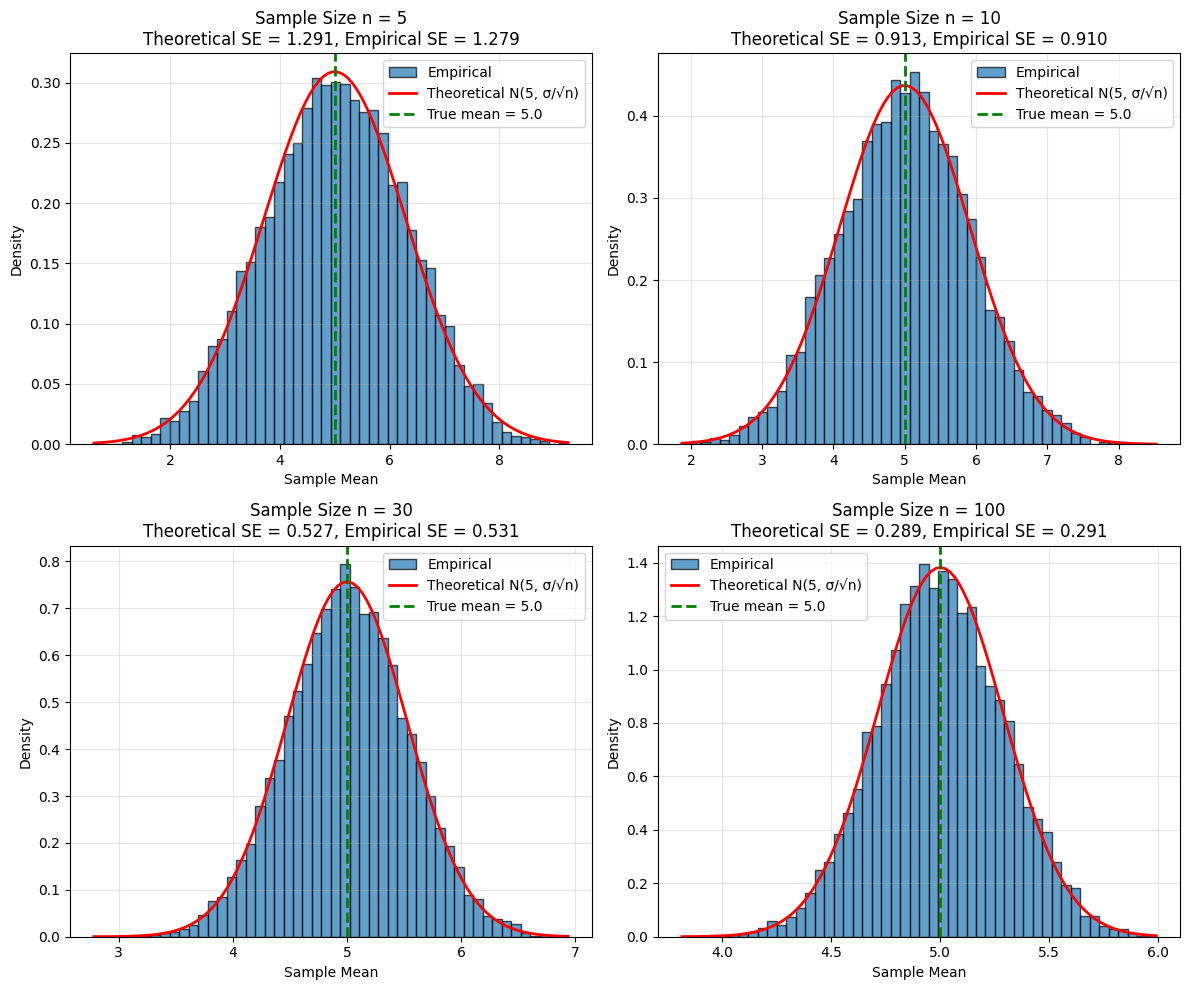
axes[idx].set_title(f'Sample Size n = {n}\nTheoretical SE = {theoretical_se:.3f}, Empirical SE = {empirical_se:.3f}')
axes[idx].legend()
axes[idx].grid(alpha=0.3)
plt.tight_layout()
plt.savefig('sampling_distribution.png', dpi=150, bbox_inches='tight')
plt.show()
print("Sample Size | Theoretical SE | Empirical SE | Ratio")
print("-" * 60)
for n in sample_sizes:
means = sample_mean(n_samples, n)
theo_se = population_std / np.sqrt(n)
emp_se = np.std(means, ddof=1)
print(f"{n:11d} | {theo_se:14.4f} | {emp_se:12.4f} | {emp_se/theo_se:5.3f}")
Sample Size | Theoretical SE | Empirical SE | Ratio
------------------------------------------------------------
5 | 1.2910 | 1.2901 | 0.999
10 | 0.9129 | 0.9205 | 1.008
30 | 0.5270 | 0.5293 | 1.004
100 | 0.2887 | 0.2885 | 0.999
Observations:
As n increases, the distribution of sample means becomes narrower (smaller SE)
The empirical SE matches the theoretical SE = σ/√n very closely
The distribution becomes more normal-looking (Central Limit Theorem!)
When Does Sampling With Replacement Work?#
The Finite Population Correction#
When sampling without replacement from a finite population of size N:
The factor \((\frac{N-n}{N-1})\) is called the finite population correction (FPC).
Rule of Thumb#
If (n < 0.05N) (sample is less than 5% of population), you can ignore the FPC:
and use \((\text{Var}(\bar{X}) = \frac{\sigma^2}{n})\).
Python Example: Finite Population#
import numpy as np
import matplotlib.pyplot as plt
# Small population
N = 100
population = np.random.normal(50, 10, N)
pop_mean = np.mean(population)
pop_var = np.var(population, ddof=0) # Population variance
# Function to sample without replacement
def sample_without_replacement(pop, sample_size, n_samples=1000):
means = []
for _ in range(n_samples):
sample = np.random.choice(pop, size=sample_size, replace=False)
means.append(np.mean(sample))
return np.array(means)
# Test different sample sizes
sample_sizes = [5, 10, 20, 40]
n_simulations = 10000
print("Sample Size | Var with FPC | Var without FPC | Empirical Var | FPC Factor")
print("-" * 80)
for n in sample_sizes:
# Sample and compute empirical variance
sample_means = sample_without_replacement(population, n, n_simulations)
empirical_var = np.var(sample_means, ddof=1)
# Theoretical variance with FPC
fpc = (N - n) / (N - 1)
var_with_fpc = (pop_var / n) * fpc
# Theoretical variance without FPC
var_without_fpc = pop_var / n
print(f"{n:11d} | {var_with_fpc:12.4f} | {var_without_fpc:15.4f} | {empirical_var:13.4f} | {fpc:10.4f}")
Sample Size | Var with FPC | Var without FPC | Empirical Var | FPC Factor
--------------------------------------------------------------------------------
5 | 20.5243 | 21.3884 | 20.0542 | 0.9596
10 | 9.7220 | 10.6942 | 9.6692 | 0.9091
20 | 4.3209 | 5.3471 | 4.3672 | 0.8081
40 | 1.6203 | 2.6736 | 1.6246 | 0.6061
Observation: When n is a significant fraction of N, the FPC makes a big difference!
Distributions Are Like Populations#
Conceptual Shift#
Instead of thinking about physical populations, we can think about probability distributions:
Population → Probability Distribution
Drawing a sample → Drawing independent samples from the distribution
Population mean μ → Expected value E[X]
Population variance σ² → Variance Var(X)
This framework allows us to work with:
Infinite populations
Repeated measurements
Stochastic processes
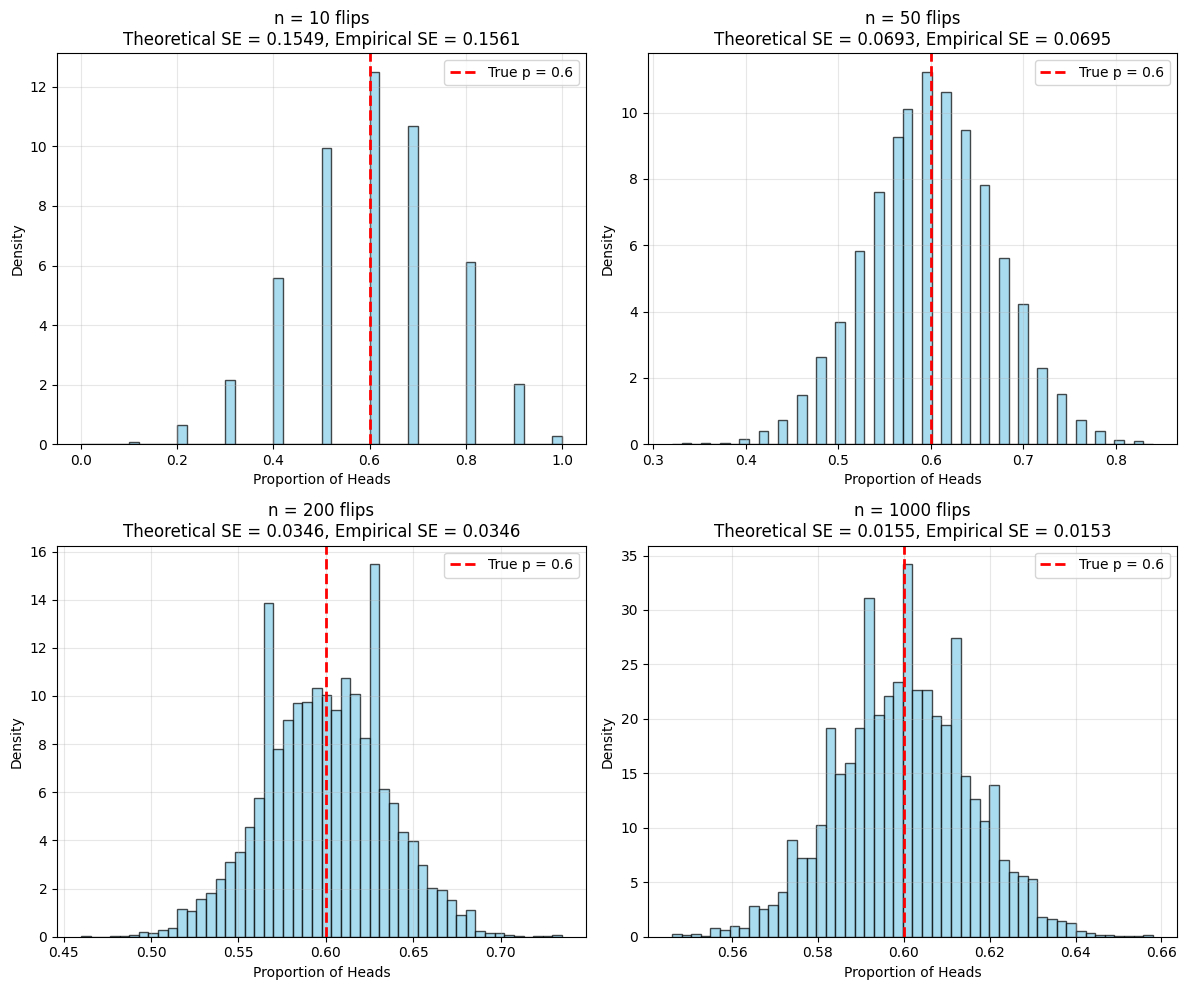
Example: Coin Flipping#
import numpy as np
import matplotlib.pyplot as plt
# "Population": Bernoulli(p=0.6) distribution
p = 0.6
pop_mean = p
pop_var = p * (1 - p)
# Flip coin n times and compute proportion of heads
def flip_experiment(p, n, n_experiments=10000):
proportions = []
for _ in range(n_experiments):
flips = np.random.binomial(1, p, n)
proportions.append(np.mean(flips))
return np.array(proportions)
# Different numbers of flips
flip_counts = [10, 50, 200, 1000]
fig, axes = plt.subplots(2, 2, figsize=(12, 10))
axes = axes.ravel()
for idx, n in enumerate(flip_counts):
props = flip_experiment(p, n)
theoretical_se = np.sqrt(pop_var / n)
empirical_se = np.std(props, ddof=1)
axes[idx].hist(props, bins=50, density=True, alpha=0.7,
edgecolor='black', color='skyblue')
axes[idx].axvline(pop_mean, color='red', linestyle='--',
linewidth=2, label=f'True p = {p}')
axes[idx].set_xlabel('Proportion of Heads')
axes[idx].set_ylabel('Density')
axes[idx].set_title(f'n = {n} flips\nTheoretical SE = {theoretical_se:.4f}, Empirical SE = {empirical_se:.4f}')
axes[idx].legend()
axes[idx].grid(alpha=0.3)
plt.tight_layout()
plt.savefig('coin_flip_sampling.png', dpi=150, bbox_inches='tight')
plt.show()